dynasor – Correlations for everyone#
dynasor is a tool for calculating total and partial dynamic structure factors as well as related correlation functions from molecular dynamics (MD) simulations. Analysis of these functions enables one to access the dynamics of a system without resorting to perturbative approaches. By combining in particular the structure factor with the cross sections (or form factors) of, e.g., neutrons, X-rays or electrons, it is also possible to directly predict experimental spectra.
Specifically dynasor can be used to calculate the following quantities:
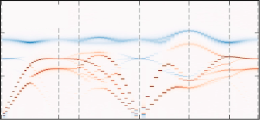
Partial and total dynamic structure factors, including the incoherent (or self) part
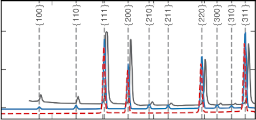
Partial and total static structure factors
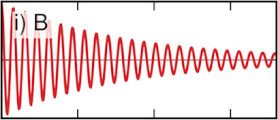
Partial and total intermediate scattering functions
Longitudinal and transverse, partial and total, current correlations
Spectral energy densities
The main input consists of a trajectory from a MD simulation, i.e., a file containing snapshots of the particle coordinates, and optionally velocities that correspond to consecutively and equally spaced points in (simulation) time. The following snippet illustrates how one can calculate dynamic structure factors from such a trajectory
traj = Trajectory('dump.xyz', trajectory_format='extxyz')
q_points = get_spherical_qpoints(traj.cell, q_max=20)
sample = compute_dynamic_structure_factors(traj, q_points=q_points, dt=5, window_size=100)
sample.write_to_npz('test.npz')
Note
For questions and help please use the dynasor discussion forum on matsci.org.
Please consult the credits page for information on how to cite dynasor.
dynasor has been developed at Chalmers University of Technology in Gothenburg, Sweden, in the Condensed Matter and Materials Theory division at the Department of Physics. dynasor and its development are hosted on gitlab.